Analysis of the makeup and functioning of microbiomes is known as microbiome analysis. The microorganisms that coexist in a specific habitat are known as a microbiome.
These microorganisms may be bacteria, viruses, fungi, protists, or archaea. The human body, the soil, and the oceans are just a few examples of the many settings where microbiomes can be found.
Despite difficulties, microbiome analysis is an effective method for determining how microbes affect human health and disease.
As this field of study develops, we are discovering more about the value of the microbiome. Moreover, we can also know how to use them to enhance human health.
Why is Microbiome analysis critical?
Microbiomes are important for several reasons. They:
- Help us digest food
- Protect us from harmful microorganisms
- Produce vitamins and other nutrients
- Regulate our immune system
- Influence our mood and behavior
- Play a role in our development and health
Microbiome analysis is a challenging task due to the following factors:
- The diversity of microorganisms in a microbiome: There are millions of different species of microorganisms, and each microbiome is unique.
- The dynamic nature of microbiomes: Microbiomes constantly change in response to factors such as diet, environment, and medication.
- The complexity of microbiome data is often vast and complex, making it challenging to analyze.
- The need for reference data: Limited reference data is available for many microbiomes. Hence making it challenging to interpret the results of microbiome analysis.
GPU Acceleration for Microbiome Analysis
GPU acceleration uses graphics processing units (GPUs), specific processors for parallel computing, to speed up complicated data analysis. Since GPUs can analyze large volumes of data. Therefore, they are ideal for microbiome analysis.
To get more information about GPU’s use in microbiology, visit Uses of GPU Computing in Microbiology
For availing GPU servers with professional-grade NVIDIA Ampere A100 | RTX A6000 | GFORCE RTX 3090 | GEFORCE RTX 1080Ti cards. Linux and Windows VPS are also available at Seimaxim.
How does it work?
Microbiome analysis can be sped up in several ways using GPUs. They can be helpful, for example:
Process large datasets
Microbiome data is frequently pervasive and diverse. It can be analyzed with challenging conventional CPUs. Specialized processors, known as GPUs, are helpful for parallel computation.
It implies that they can do several calculations simultaneously. It qualifies them for jobs like microbiome analysis, which may require data processing.
One microbiome sample, for instance, may contain millions of DNA sequences. Researchers need to employ several computer techniques to analyze these sequences.
On conventional CPUs, these techniques can be computationally expensive and time-consuming. However, GPU technology can accelerate these techniques by orders of magnitude.
GPUs can perform multiple calculations at the same time. For example, a single GPU can process thousands of DNA sequences simultaneously.
It can be helpful to analyze large datasets of microbiome data much faster than traditional CPUs.
In addition to speeding up the analysis process, GPUs can also improve the accuracy of microbiome analysis.
GPUs can reduce the errors introduced during the analysis process. Moreover, GPUs are more precise than CPUs when performing calculations.
Run complex algorithms
To analyze the microbiome, some challenging algorithms that can be run on GPUs include:
Alignment algorithms
DNA sequence comparisons are made using alignment methods. Although this is a computationally intensive job, GPUs can significantly speed it up.
Machine learning algorithms
Algorithms for machine learning are employed to find patterns and trends in data.
Although it is an effective tool for microbiome investigation, running it on conventional CPUs might be computationally expensive. Machine learning algorithms for microbiome analysis can be sped up using GPUs.
Statistical analysis algorithms
Algorithms for statistical analysis are helpful to examine data and derive conclusions from it.
Although it is a typical activity in microbiome analysis, running it on conventional CPUs might be costly in terms of processing costs. Algorithms for statistical analysis of the microbiome can be sped up using GPUs.
Visualize data
There are many different ways to visualize microbiome data. Researchers can use this to find patterns and trends in the data.
However, it might be computationally expensive to visualize microbiome data, particularly for big datasets.
Its simultaneous computation capabilities can accelerate microbiome data visualization with GPUs.
Therefore, it indicates they are substantially faster than conventional CPUs at visualizing significant amounts of microbiome data.
For availing GPU servers with professional-grade NVIDIA Ampere A100 | RTX A6000 | GFORCE RTX 3090 | GEFORCE RTX 1080Ti cards. Linux and Windows VPS are also available at Seimaxim. Some of the ways that GPUs can be used to visualize microbiome data include:
Heatmaps
A heatmap is a visualization that shows the number of microbes in a microbiome sample. Each cell in the two-dimensional matrix reflects the quantity of a specific type of microbe in a particular area.
Each cell’s color indicates the number of microorganisms. Therefore, deeper hues denote more numbers, and lighter shades represent fewer numbers. GPUs generate heatmaps of microbiome data by performing the following steps:
- The microbiome data is divided into a grid of cells.
- The abundance of each microorganism in each cell is calculated.
- The color of each cell is assigned based on the quantity of the microorganism in that cell.
A GPU can divide the microbiome data into a grid of cells in the first stage. Therefore, GPUs’ ability to perform parallel computing enables them to split the work of organizing data into a grid of cells into numerous more minor tasks. Then, carry out each task simultaneously.
This can substantially speed up the data partitioning into a grid of cells.
A GPU is beneficial for the second stage, which involves figuring out how many microorganisms are present in each cell. All of this is because of how adept GPUs handle mathematical operations.
Therefore, they can substantially speed up the procedure of estimating the amount of each microorganism in each cell because they can carry out these computations much more quickly than CPUs.
A GPU can also be helpful for the third stage, which determines the color of each cell based on how many microorganisms are present in that particular cell. GPUs can perform parallel computing.
Therefore enabling them to assign the colors to the cells concurrently. It substantially hastens the process of giving the cells their respective colors.
Heatmap of microbial communities in wastewater in winters
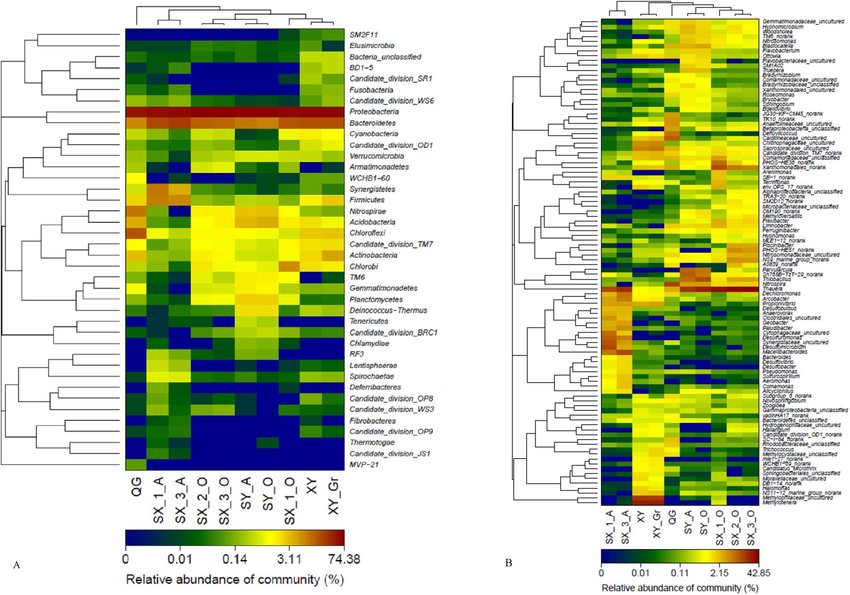
The distinct microbial communities that are present in the soil sample might be identified using this heatmap.
Additionally, it might be used to monitor alterations in the microbial community over time or in response to various environmental factors.
Tree
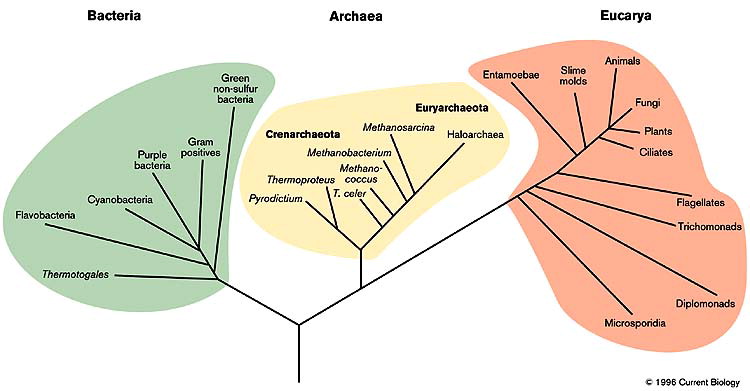
We can visually observe the relationships between the microorganisms in a microbiome sample using trees.
Therefore, they are often helpful in distinguishing between the various microbial communities present in a sample.
Moreover, they are also helpful in monitoring changes in the microbial community over time or in response to different environmental circumstances.
Microbiome data trees can be created on GPUs significantly more quickly than on CPUs.
This is because GPUs can perform parallel computing, which allows them to split the process of creating the tree into numerous more minor tasks and then carry out each task simultaneously.
It can result in a significant speedup in the tree generation process.
Various methods can produce the generation of microbiome data trees on GPUs. Utilizing a hierarchical clustering technique is one typical strategy.
Similar microorganisms are grouped by hierarchical clustering methods to function. GPUs can expedite the hierarchical clustering process by doing the clustering algorithms in parallel.
Employing a phylogenetic tree algorithm is another typical method for creating trees from microbiome data.
Therefore, reconstructing the evolutionary links of microorganisms is how phylogenetic tree algorithms function.
GPUs can accelerate the phylogenetic tree reconstruction procedure by running the tree reconstruction calculations in parallel.
GPUs can also be helpful to display microbiome data as trees. Researchers who desire to delve deeper into the interactions between microorganisms may find this useful.
Moreover, Researchers can study the data in real-time using interactive visualizations of microbiome data trees produced by GPUs.
A group of researchers at the University of California, San Diego, used GPUs to generate trees of microbiome data from the human gut. The researchers found that GPUs could generate trees of microbiome data much faster than CPUs.
This enables them to study the relationships between microorganisms in the human gut in more detail.
Networks
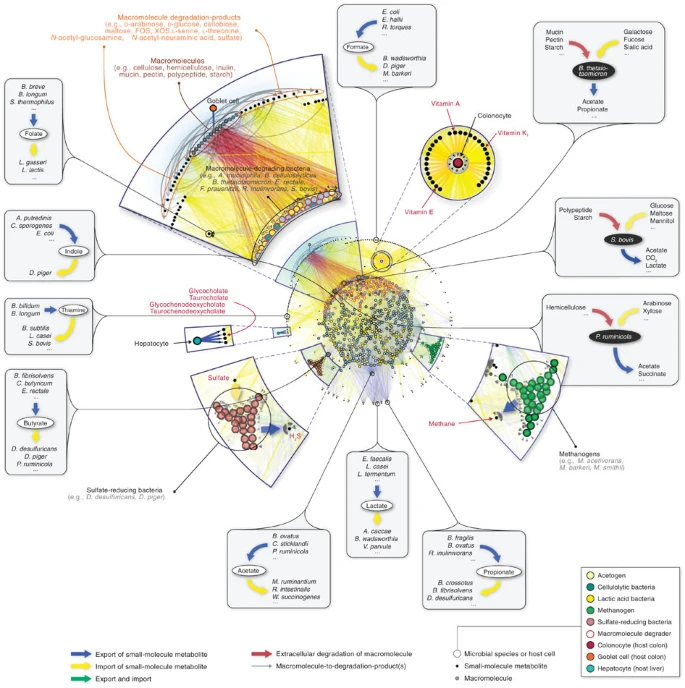
Networks are helpful in visualization to show how different microorganisms interact in a microbiome sample.
Therefore, they are frequently employed to distinguish between the microbial communities in a sample and monitor microbial community changes over time or in response to various environmental factors.
GPUs can be helpful in various ways to create networks using microbiome data. A correlation network algorithm is one such method.
Correlation network methods are used to find pairs of microorganisms that are highly associated with one another.
Therefore, GPUs can hasten the development of correlation networks by doing the correlation calculations in parallel.
Applying a metabolic network algorithm is another typical method for creating networks of microbiome data.
Algorithms for metabolic networks detect pairs of microorganisms that collaborate on a metabolic pathway.
However, by running the route analysis calculations in parallel, GPUs can hasten the creation of metabolic networks.
Some real-time examples of using GPU for microbiome analysis
Here are some examples of GPU-accelerated microbiome studies
- A study by researchers at the University of California, San Diego, used GPU acceleration to study the human gut microbiome in people with type 2 diabetes. The researchers found that people with type 2 diabetes had a different microbiome composition than people without type 2 diabetes. It suggests that the microbiome may play a role in developing type 2 diabetes.
- Another study by researchers at the University of Texas Southwestern Medical Center used GPU acceleration to study the soil microbiome in different agricultural fields. The researchers found that the microbiome composition of the soil varied depending on the type of agriculture practiced. The microbiome may play a role in the health of agricultural soils.
- A study by researchers at the University of California, Berkeley, used GPU acceleration to study the human skin microbiome in people with acne. The researchers found that people with acne had a different microbiome composition than people without acne. The microbiome may play a role in the development of acne.
- Another study by researchers at the University of Cambridge used GPU acceleration to study the microbiome of the human mouth in people with gum disease. The researchers found that people with gum disease had a different microbiome composition than people without gum disease. The microbiome may play a role in the development of gum disease.
Best practices for GPU-accelerated microbiome analysis
Choosing the right GPU
Compute capacity
A GPU’s performance can be gauged by its computing capacity. So, GPU-accelerated microbiome analysis will go faster on GPUs with more computing power.
Memory
GPUs require a lot of memory to store the microbiome data and the research results. GPUs can process larger datasets with more RAM.
Price
GPUs have a range in price. Therefore, Picking a GPU that meets your requirements is critical while remaining within your price range.
For availing GPU servers with professional-grade NVIDIA Ampere A100 | RTX A6000 | GFORCE RTX 3090 | GEFORCE RTX 1080Ti cards. Linux and Windows VPS are also available at Seimaxim.
Using specialized software
For GPU-accelerated microbiome analysis, there are several specific software programs available. These software programs have been created to make the most of the unique features of GPUs.
The following aspects should be kept into account when selecting specialized software:Features: The software bundle must provide the required elements for your investigation.
Usability: The software program should be simple, even if you are not a pro in GPU programming.
Support: The authors of the software product should provide excellent support.
Standardizing data formats
A hurdle for GPU-accelerated microbiome analysis, as was before highlighted, is the need for standardized data formats. Standardizing data formats for storing microbiome data is one approach to solving this problem.
Numerous groups are striving to create standardized data formats for microbiome data. These organizations include the Global Alliance for Microbiome Standards (GAMeS) and the Microbiome Standards Institute (MSI).
We can simplify the usage of GPU acceleration tools by researchers by standardizing the data formats for storing microbiome data. Microbiome analysis will become quicker and more effective as a result.
Additional best practices
Here are some additional best practices for GPU-accelerated microbiome analysis:
- Optimize your code: When using GPU acceleration software, it is crucial to optimize your code to take advantage of the specific capabilities of GPUs.
- Use appropriate batch sizes: Batching is a technique that can help improve the performance of GPU-accelerated microbiome analysis. Therefore, when batching data, you group similar data together and then process the batch as a single unit. It can improve performance because GPUs can process multiple data points simultaneously.
- Use a GPU cluster: If you can access a GPU cluster, you can use it to perform GPU-accelerated microbiome analysis on even larger datasets. GPU clusters are made up of multiple GPUs that are connected. Hence, It allows you to distribute the workload of the study across multiple GPUs.
Challenges of GPU Acceleration for Microbiome Analysis
The need for specialized software
The requirement for specialized software makes GPU acceleration for microbiome investigation difficult.
Therefore, most tools for general-purpose microbiome investigation do not support GPU acceleration.
Moreover, This implies that scientists must employ specialized software created especially for GPU acceleration.
The creation of specialized software for GPU acceleration comes with several difficulties.
The fact that GPUs and CPUs differ significantly creates a hurdle.
This means that to create software that can use GPUs’ capabilities, software developers must be conversant with the unique architecture of those devices.
Another improvement area is that integrating GPU acceleration can be difficult.
This is because GPUs are built for parallel processing, enabling them to carry out multiple calculations simultaneously.
Also, software developers must carefully plan their programs to maximize this parallel processing potential.
The high cost of GPUs
The high price of GPUs adds another difficulty for GPU acceleration in microbiome investigation.
Compared to CPUs, GPUs are substantially more expensive. Therefore, It can make using GPU acceleration challenging for researchers on a tight budget.
Numerous reasons contribute to the high price of GPUs.
One of the reasons is that it takes more effort to build GPUs than CPUs. Another factor is the rising demand for GPUs across various sectors, including gaming and artificial intelligence.
The lack of standardized data formats
The absence of standardized data formats is the third obstacle to GPU acceleration for microbiome analysis.
Hence, different formats can be helpful to collect microbiome data; not all are supported by GPU acceleration software
Researchers may find it challenging to exploit GPU acceleration due to the need for standardized data formats.
It takes time for researchers to transfer their data into a GPU acceleration software-compatible format.
Conclusion
The high price of GPUs adds another difficulty for GPU acceleration in microbiome investigation.
Compared to CPUs, GPUs are substantially more expensive. It can make using GPU acceleration challenging for researchers on a tight budget.
Numerous reasons contribute to the high price of GPUs. One of the reasons is that it takes more effort to build GPUs than CPUs.
Another factor is the rising demand for GPUs across various sectors, including gaming and artificial intelligence.
These innovations could improve the effectiveness and precision of microbiome analysis.
This could result in a fresh understanding of how the microbiome affects a variety of illnesses, such as cancer, diabetes, and obesity.
Overall, GPU-accelerated microbiome analysis has a promising future. As technology advances, it is expected to be utilized more frequently and provide fresh insights into the human microbiome.